What is the difference between the ERCCs and the SIRVs?
ERCC Spike-In Controls (ERCCs, Ambion, Thermo Fisher) are a set of 92 synthetic, polyadenylated RNAs of varying length (250 - 2000nt), GC content, and concentration. They enable assessment of dynamic range, dose response, lower limit of detection, and fold-change response in RNA sequencing pipelines. However, because ERCCs are mono-exonic, they cannot be used to measure or validate splice variants - one of the main challenges when sequencing complex transcriptomes.
In contrast, Lexogen´s SIRV controls (Spike-In Transcript Variants) consist of 69 synthetic, polyadenylated isoforms designed specifically to evaluate isoform detection and quantification in RNA sequencing workflows. By providing a well-defined “ground truth”, SIRVs allow for robust comparison of experiments, enabling results from a controlled subset of spike-in reads to be extrapolated to the broader sample. Similar to ERCs, SIRVs also support the assessment of dynamic range, dose response, detection limits, and fold-change response within the context of transcript variant detection.
Additionally, the long SIRV controls address the need for spike-in transcripts exceeding 2.5 kb. Spanning 4 - 12kb, these long transcripts are particularly valuable for validating long-read sequencing platforms and RNA-Seq setups.
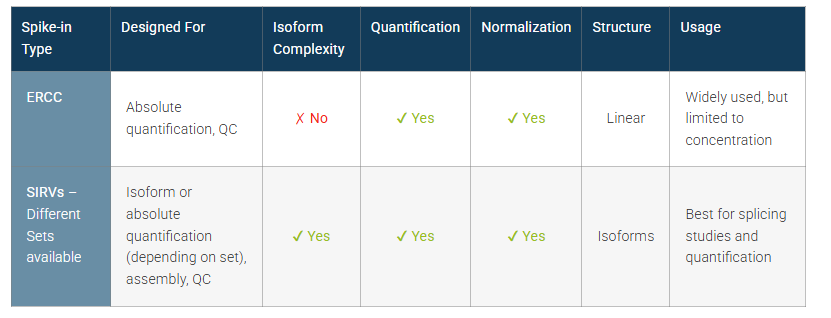
Table 1 | Spike-in RNA Controls for gene expression, Quantification and Isoform Detection
