How many PCR cycles are needed to amplify QuantSeq libraries?
The optimal number of PCR cycles for a given input amount of total RNA can vary by up to four cycles, depending on sample quality and origin. The optimal cycle number for your specific sample type should be determined using the qPCR assay.
The table below shows Endpoint PCR cycle numbers for QuantSeq FWD libraries prepared from Universal Human Reference RNA (UHRR) input amounts. This should be taken as a guideline only! Optimal cycle numbers could exceed these ranges depending on the sample type and quality (i.e., species, tissue, RNA quality e.g., FFPE RNA, etc.).
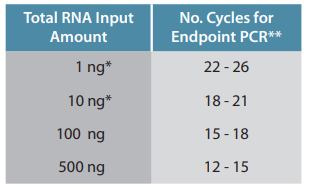
*Using low input protocol modifications with 1 hour incubation at 42°C (Step 4- Appendix B, p19).
**These values are provided as reference only! Sample type influences the optimal cycle number, which should be determined by qPCR assay.
qPCR Assay for Endpoint PCR Cycle Number Determination
The qPCR assay to determine optimal PCR cycle numbers for library amplification is performed using an aliquot of purified double-stranded cDNA, prior to the final library amplification.
To perform the qPCR assay you will need:
The PCR Add-on and Reamplification Kit V2 (Cat. No. 208), and
SYBR Green I nucleic acid stain (10,000X in DMSO, not provided in the kit. We recommend using Sigma-Aldrich, Cat. No. S9430 or ThermoFisher, Cat. No. S7585)
ATTENTION! The use of SYBR Green I-containing qPCR mastermixes from other vendors is not recommended.
Protocol
The recommended protocol for the qPCR assay is provided in the PCR Add-on and Reamplification Kit V2 User Guide, as well as in the Appendices of each QuantSeq 3' mRNA-Seq Library Prep Kit User Guide.
Additional Resources
For additional information on the qPCR Assay consult the following online FAQs:
https://faqs.lexogen.com/faq/what-do-i-need-for-the-qpcr-assay
https://faqs.lexogen.com/faq/how-do-i-calculate-endpoint-cycle-numbers
